
American scientists planned to work with the Wuhan Institute of Virology to engineer novel coronaviruses with the features of SARS-CoV-2 the year before the virus emerged from that city, according to documents obtained by U.S. Right to Know.
While rare in nature, these features were central to the esoteric research interests of the scientists working with the Wuhan lab, those documents show.
Scientists divided over the so-called “lab leak” and natural origin hypotheses have for years pored over the arcane language of a U.S.-China research proposal called “DEFUSE” describing coronavirus engineering experiments.
The DEFUSE grant proposal was led by EcoHealth Alliance President Peter Daszak.
Now, drafts and notes uncovered through the Freedom of Information Act reveal fresh details about the intended research.
Specifically, the scientists sought to insert furin cleavage sites at the S1/S2 junction of the spike protein; to assemble synthetic viruses in six segments; to identify coronaviruses up to 25 percent different from SARS; and to select for receptor binding domains adept at infecting human receptors.
The genome of SARS-CoV-2, the virus that causes COVID-19, matches the viruses described in the research proposal:
- SARS-CoV-2 has a furin cleavage site positioned in the spike protein at the S1/S2 junction. The furin cleavage site supercharged the virus into the worst pandemic pathogen in a century. Virologists have yet to identify one in any other related coronavirus.
- SARS-CoV-2 can be divided into six contiguous genomic pieces by the restriction enzymes Bsal and BsmBI. These restriction enzymes occur in nature but can also be used in the lab to splice viruses. A trio of scientists estimated in a 2022 analysis that the likelihood of seeing the pattern found in SARS-CoV-2 in nature would be remote. Orders for one of these restriction enzymes, BsmBI, can be found in the documents.
- SARS-CoV-2 emerged highly infectious without evolving much in humans. The virus “came out of the box ready to infect.” The receptor binding domain appeared “finely tuned” for the human ACE2 receptor, yet had little genetic variation when first spilling over into humans, presenting a difficult “paradox” to virologists who sought to prove it emerged naturally. The documents confirm the scientists working with the Wuhan lab sought to select for receptor binding domains that bind well to human ACE2 in their research.
- The genome of SARS-CoV-2 falls within the range of a 25 percent genetic difference from SARS.
The documents reveal for the first time that a virologist working with the Wuhan lab planned to engineer new spike proteins – in contrast with the collaboration’s public work to insert whole spike proteins into viral backbones. Language in the proposal indicates this work may have involved unpublished viruses, generating unpublished engineered spike proteins.
This American virologist, University of North Carolina Prof. Ralph Baric, was set to engineer twenty or more “chimeric” SARS-related viral spike proteins per year of the proposal, and two to five full-length engineered SARS-related viruses. Documents previously reported by U.S. Right to Know show that some of the experimentation could secretly occur in Wuhan at a lower biosafety level than specified in the grant, apparently to save costs.
The documents challenge an argument made by the National Institutes of Health and some virologists against the relevance of the research proposal to the origins of the pandemic. They have argued that this U.S.-China scientific collaboration only planned to engineer viruses starting with viral backbones already in the public literature, and that these viral backbones are too dissimilar to have played a role in the pandemic.
The new documents however reveal that the scientists planned to use new reverse genetics systems and test viruses in vivo — in other words, to engineer live viruses with novel backbones.
The documents describe the SARS-related viruses to be studied in the grant as posing “a clear-and-present danger of a new SARS-like pandemic.”
The documents do not prove a precise step-by-step instruction manual for how SARS-CoV-2 was generated in the lab. The genomes of some of the SARS-related viruses the scientists planned to work with remain unknown. But they do describe experiments that could have generated the virus’ rare properties. They detail the scientists’ interest in working with viruses precisely like SARS-CoV-2.
The grant proposal was submitted to the Defense Advanced Research Projects Agency, which rejected the project. Whether the research was funded through other means remains unknown. Baric had engineered unknown spike proteins by the time the proposal was submitted.
Nonetheless, the documents suggest that some of the data central to the worst pandemic in a century may be found not only in China, but also in the U.S.
“When the Wuhan Institute of Virology published their first paper on the pandemic virus, they made no mention of the unique furin cleavage site despite having recently authored DEFUSE, declaring that they were on the lookout for these concerning features in novel SARS-like viruses,” Broad Institute molecular biologist Alina Chan said. “We need to get all of the exchanges between the Wuhan Institute of Virology and its US collaborators in 2018 and especially 2019 – the year of the pandemic.”
Furin cleavage site at the S1/S2 boundary
While the language of the formal DEFUSE proposal called for the insertion of “human specific proteolytic cleavage sites” in a portion of the spike protein called the “S2′,” earlier drafts of DEFUSE were more explicit.
Proteolytic cleavage sites is a more general term than furin cleavage sites. It refers to a combination of amino acids that allow enzymes to cleave the spike protein, which helps viruses like SARS-CoV-2 enter human cells. But proteolytic cleavage sites can be activated by a variety of enzymes, not just furin.
This language in the final, formal proposal has been a sticking point in the debates over the DEFUSE documents.
Scripps Institute virologist Kristian Andersen — an advocate for the natural origin theory — has argued that DEFUSE holds little relevance to the origins of COVID because the grant calls for the introduction of proteolytic cleavage sites at the S2′ position of the spike protein.
However the earlier drafts show the scientists’ particular interest in furin cleavage sites. The documents suggest the scientists were not yet certain about the relative importance of the other cleavage sites.
Earlier drafts also precisely specify the positions of the intended insertions, including ones that correspond to the S1/S2 junction, not just at S2′.
“Tissue culture adaptations sometimes introduce a furin cleavage site which can direct entry processes, usually by cleaving S at positions 757 and 900 in S2′ of other CoV, but not SARS,” the grant reads.
The position 900 is the S2′ site, and position 757 is the S1/S2 site.
A comment on the proposal clarifies that this language cites Baric’s “Figure C.”
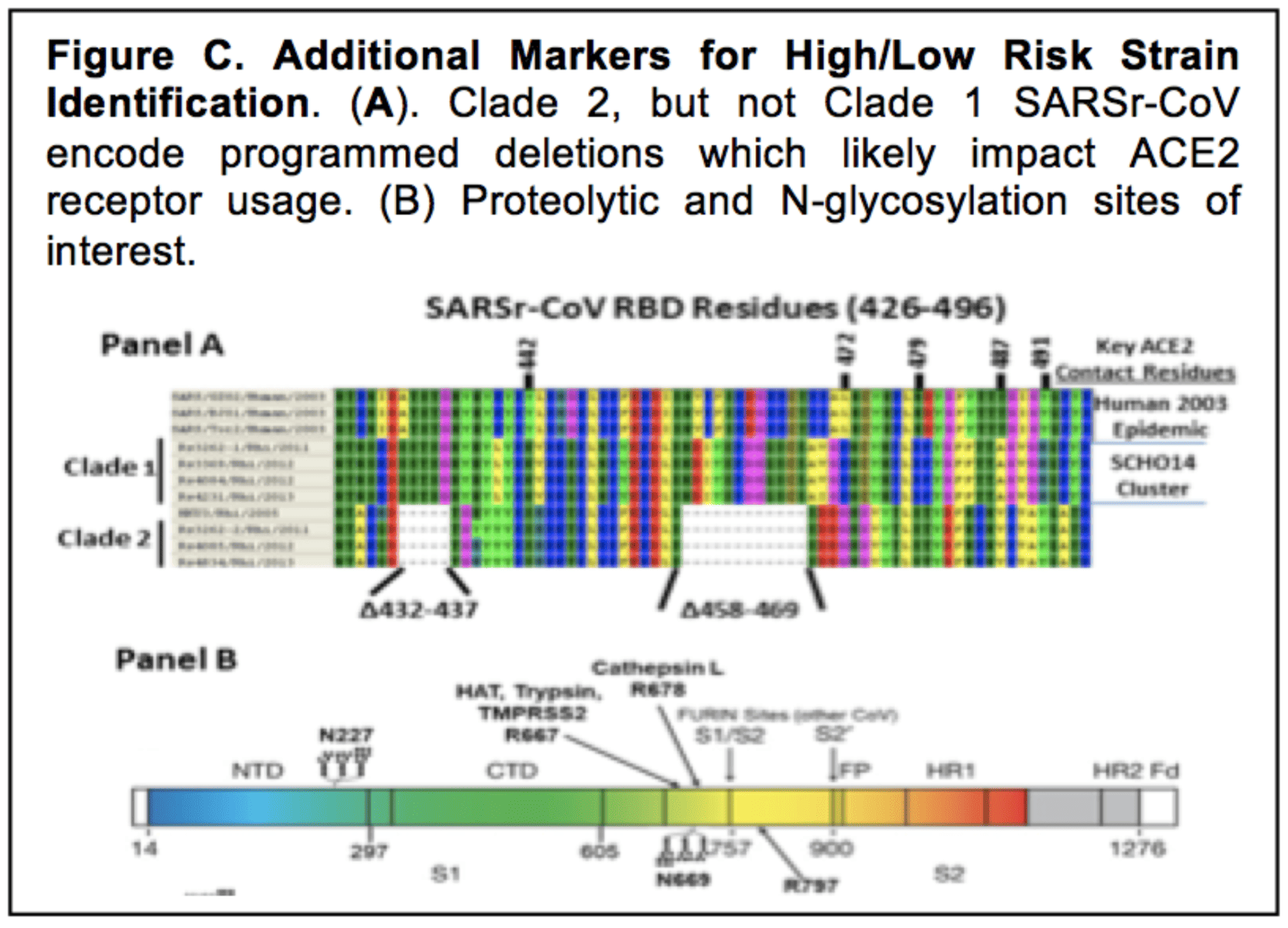
Figure C shows a furin cleavage site at the S1/S2 boundary, precisely where the furin cleavage site in SARS-CoV-2 is found.
The documents suggest the research group had identified a way to isolate SARS-related coronaviruses with cleavage sites or to insert them, some scientists say.
Restriction sites
Some scientists who favor the natural origin theory have argued that the Wuhan lab would have only employed familiar backbones in the published literature and swapped out spike proteins. Because these backbones in the published literature are too genetically dissimilar to have generated SARS-CoV-2, they have argued the DEFUSE proposal is irrelevant to the pandemic.
However, more candid early drafts of the grant show the researchers planned to test engineered spike proteins in these familiar backbones as an initial test that would help them prioritize genomes for the next step: the generation of synthetic viruses in six pieces.
The spike proteins identified by the group this way to have “pre-epidemic potential” would be employed in the next step, the generation of “full genome length viable viruses.”
The documents show that the scientists behind DEFUSE proposed a strategy to stitch SARS-related viral genomes together using six pieces.
They planned to assemble full-length synthetic viruses. These viruses were to be assembled from consensus sequences — sequences that summarize the most common base pairs among a group of closely related viruses.
These viruses could have up to a 5 percent nucleotide variation from one another. RaTG13, one of SARS-CoV-2 closest cousin viruses, which was sequenced by the Wuhan Institute of Virology, is 4 percent different than SARS-CoV-2.
“We will identify the best consensus candidate and synthesize the genome using commercial vendors (e.g., BioBasic, etc.), as six contiguous cDNA pieces linked by unique restriction endonuclease sites that do not disturb the coding sequence, but allow for full length genome assembly,” the grant states.
The documents show they anticipated synthesizing viruses to be cheap.
The researchers planned to infect mice with humanized lung cells with these full-length synthetic viruses.
This language in the newly revealed documents echoes a 2022 analysis that uncovered a pattern of two restriction enzymes, BsmBI and BasI, that segmented the SARS-CoV-2 viral genome into six even pieces. The scientists estimated the likelihood of observing this pattern of evenly spaced segments in nature to be highly improbable.
At the time, it was dismissed by Andersen as “kindergarten molecular biology.”
“Many virologists said that our analysis was flawed, claiming that the aim must have been to replace the entire spike … or that similar patterns can be found in virtually any coronavirus genome,” said University of Wuerzburg molecular immunologist Valentin Bruttel, a coauthor of that analysis. “Exactly as we had postulated, they planned to use 6 segments to assemble synthetic viruses.”
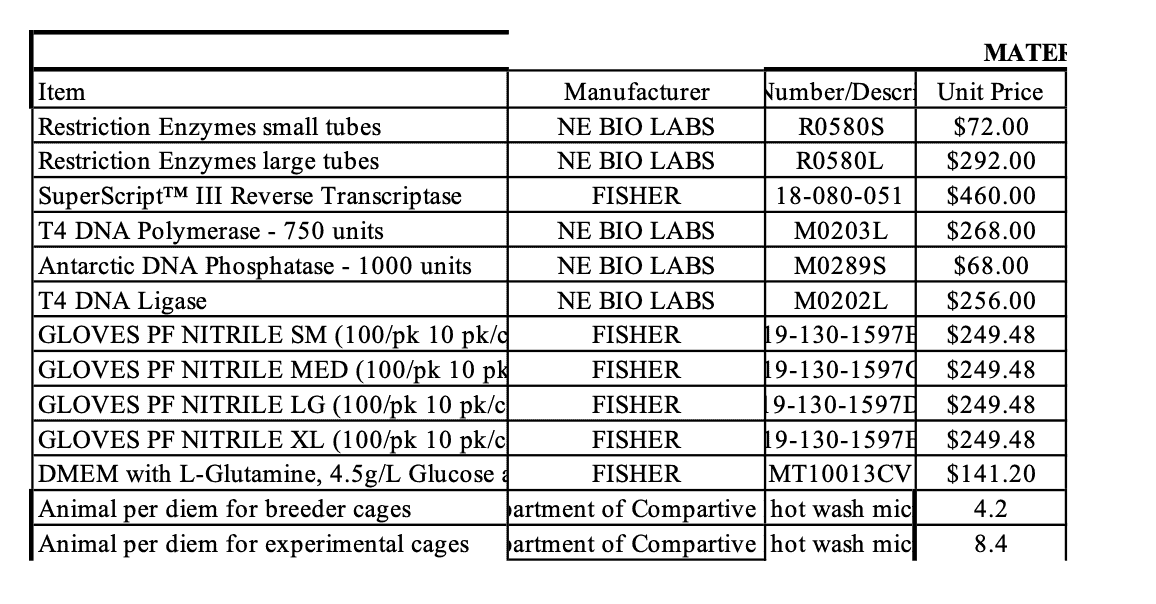
The newly revealed documents also include an order from New England Biolabs for BsmBI, one of these key restriction enzymes. They also planned to buy other unspecified restriction enzymes. New England Biolabs also sells BsaI restriction enzymes, in addition to hundreds of others.
In the ordering of the documents obtained by U.S. Right to Know, this budget table appears right after an email in which Tonie Rocke, a USGS collaborator on the project, says her budget is attached. While Rocke was set to collaborate with Baric, it’s not clear that she was central to this genetic engineering work.
Still, some scientists say this finding is akin to a “smoking gun” in favor of the lab hypothesis.
But the finding is likely to stir debate.
Some of the same restriction enzyme sites identified as possible signals in SARS-CoV-2 have been identified in viruses closely related to SARS-CoV-2 in nature, indicating they could have resulted from recombination, not engineering.
The receptor binding domain
Newly available notes from calls related to the DEFUSE proposal show that the research group was interested in SARS-related coronaviruses that resemble SARS-CoV-2 in a pivotal portion of the viral genome that attaches to human cells — the receptor binding domain.
Baric planned to screen for receptor binding domains with epidemic potential by looking for cleavage sites and receptor binding domains adept at attaching to human ACE2 receptors.
In addition to the furin cleavage site, the almost immediate ability of SARS-CoV-2 to spread among humans without having to evolve much – the fact it was “well adapted” – has been a red flag for a possible lab origin since the early months of the pandemic.
The notes also show that Baric engineered spike proteins that do not appear in the public scientific literature, and that this work may have already been underway as the proposal was submitted to DARPA.
“RB [Ralph Baric] has already generated SARS-like chimeras w/ RBD [receptor binding domain] from group of bat viruses called 293 (for S1) which is 20% different than epidemic strains, and S2 region from HK3 which is 20% diff,” the notes read, apparently referring to engineered spike proteins generated from two different strains of bat viruses.
While HKU3 bat viruses are known, the reference to “bat viruses called 293” is ambiguous, and does not appear to refer to any public group of viruses.
Novel viruses at the WIV
Many of the viruses sampled by the Wuhan lab were not evaluated for transmissibility or pathogenicity at the time the grant proposal was submitted. Their genomes may not be public.
“The Wuhan Institute of Virology team will continue to collect biodiversity surveys from SARSr-CoV viruses of bat caves across S. China,” the documents read. “They have large, but incomplete collections of SARSr-CoVs sequences, most of which have not been evaluated for pre-epidemic potential.”
Earlier versions of the grant proposal identify Wuhan Institute of Virology Senior Scientist Shi Zhengli’s lab as conducting experimental tests, though this was eventually concealed from DARPA.
The documents show the research group had a hotspot for sampling bat viruses in Laos, in addition to their better known sampling efforts in Southern China.
The scientists also describe their research aims as including protecting U.S. soldiers stationed in Southeast Asia. Some of the closest cousin viruses to SARS-CoV-2, including a virus called Banal-20-52, were identified in Laos.
U.S. Right to Know obtained the documents reported on for this story from a Freedom of Information Act request to the United States Geological Survey. More documents and reporting on Covid origins is available on our website.









